METACHROMATIC STAINING
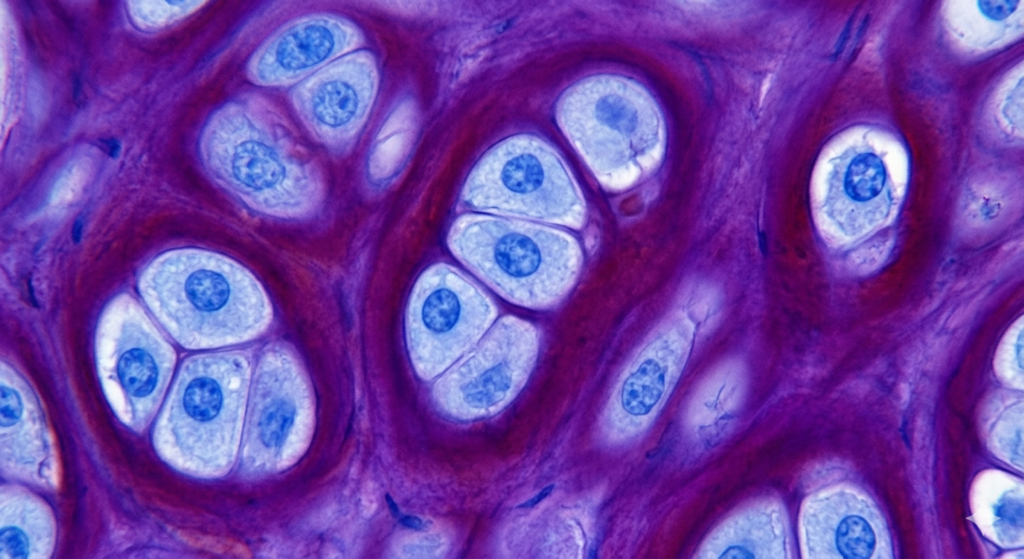
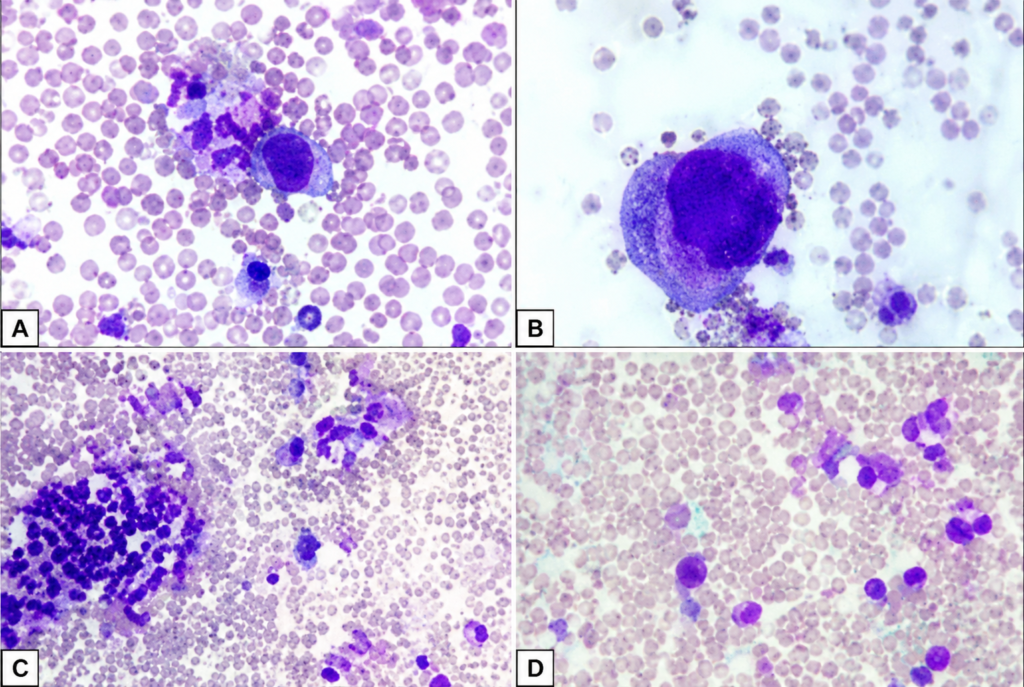
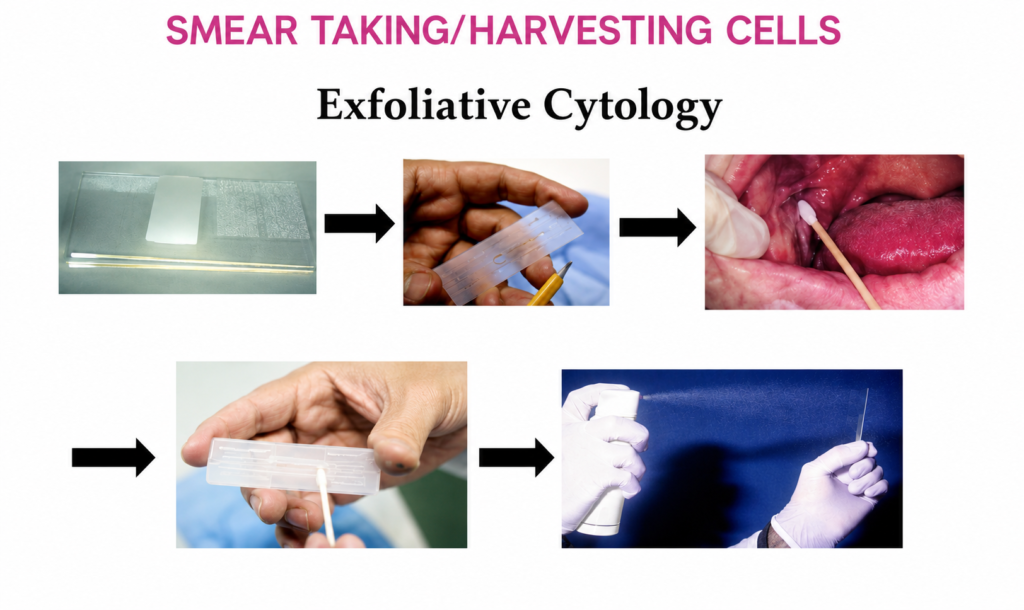
INTRODUCTION:- There are certain basic dyes belonging to aniline group that will differentiate particular tissue components by giving them a different color to that of original dye. The phenomenon is known as metachromasia. METACHROMASIA:-Metachromasia takes place when certain negatively charged groups on the tissue react with cationic dyes. On polymerization the original colour of the dye changes to another colour (eg mast cell stain pink with toluidine blue). Thionin and toluidine blue dyes are commonly used for quick staining of frozen selection using their metachromatic property to stain nucleus and cytoplasm differently.Metachromasia is enhanced when intermolecular distances are reduced. Factors which enhance metachromasia are:- 1. Increasing concentration of dye. 2. Decreasing temperature. 3. pH 4. Water a polar solvent, contributes to the efficiency of van der Waal’s forces by which the molecules are held together. In tissues, where there is a high concentration of anions e.g. in sulphated mucopolysaccharides, the cationic dye molecules may be held in such close proximity to one another that van der Waal’s forces can exert their influence and cause the dye to polymerize. Consequently the colour changes from blue to red. Tissue components often demonstrated by metachromatic stains:- Amyloid material, Mast cell granules Mucin Cartilage Amyloid Stain -Various stains are used to demonstrate amyloid CRYSTAL VIOLET STAIN FOR AMYLOID:- Aim: To demonstrate amyloid in tissue sections. Principle: Amyloid (a glycoprotein) exhibits metachromasia in tissue sections when stained with crystal violet and other cationic dyes. Control: positive control. Reagents:- Crystal violet solution Stock solution Crystal violet 14 gm 95% alcohol 100 ml Working solution Stock solution 10 ml Distilled water 300 ml Concentrated hydrochloric acid 1 ml Procedure:- *Deparaffinize and bring the sections to water. *Put working crystal violet solution for 1 to 2 minutes and check under microscope. *Rinse in tap water. *Mount in water or in water soluble media. *Put on the coverslip seal the edges with nail polish (Do not let it dry.) Result:- Amyloid purple violet Other tissues blue CONGO-RED STAIN FOR AMYLOID:- Aim:- To demontrate amyloid in tissues. Principle:- Diazo dye attaches itself to amyloid fibrils. The union is affected by H bonds between the OH groups of amyloid and amino side groups of the dye. Congo red dye forms non-polar hydrogen bonds with amyloid. The green birefringence of congo red stained amyloid by polarized light is considered diagnostic of amyloid. Control:- Known positive tissue. Reagents Congo red solution Congo red 1.0gm Distilled water 100ml Saturated solution of Lithium Carbonate Lithium carbonate 1.3gm Distilled water 100ml Procedure:- *Bring section to water. *Pour congo red solution for 20 minutes. *Pour off the solution and cover the slide with lithium carbonate for 1.5 minutes to differentiate. *Wash with water. *Counter-stain with hematoxyline for 5 minutes. *Differentiate with 1% acid alcohol. *Wash in running tap water. *Dehydrate, clear in xylene and mount in DPX. Result:- Amyloid bright red which gives apple green birefringence in polarized light. Nuclei blue Other structures unstained to yellow Notes:- 1. Sections must be cut at 8 to 10 microns for birefringence 2. Solution must be filtered through glass wool, not paper filters for birefringence to occur 3. Tissue fixed in solutions other than formalin may display false positive birefringence.